n=arg or fsmethod=arg (default=“map”) | Number of factors: “kaiser” (Kaiser-Guttman greater than mean), “mineigen” (Minimum eigenvalue criterion; specified using “eiglimit”), “varfrac” (fraction of variance accounted for; specified using “varlimit”), “map” (Velicer’s Minimum Average Partial method), “bstick” (comparison with broken stick distribution), “parallel” (parallel analysis: number of replications specified using “pnreps”; “pquant” indicates the quantile method value if employed), “scree” (standard error scree method), “bn” (Bai and Ng (2002)), “ah” (Ahn and Horenstein (2013)), integer (user-specified integer value). |
eiglimit=number (default=1) | Limit value for retaining factors using the eigenvalue comparison (where “n=mineigen”). |
varlimit=number (default=0.5) | Fraction of total variance explained limit for retaining factors using the variance limit criterion (where “n=varlimit”). |
porig | Use the unreduced matrix for parallel analysis (the default is to use the reduced matrix). For parallel analysis only (“n=parallel”). |
preps= integer (default=100) | Number of parallel analysis repetitions. For parallel analysis only (“n=parallel”). |
pquant=number | Quantile value for parallel analysis comparison (if not specified, the mean value will be employed). For parallel analysis only (“n=parallel”). |
pseed=positive integer | Seed the random number generator for parallel analysis. If not specified, EViews will seed the random number generator with a single integer draw from the default global random number generator. For parallel analysis only (“n=parallel”). |
Type of random number generator for the simulation: improved Knuth generator (“kn”), improved Mersenne Twister (“mt”), Knuth’s (1997) lagged Fibonacci generator used in EViews 4 (“kn4”) L’Ecuyer’s (1999) combined multiple recursive generator (“le”), Matsumoto and Nishimura’s (1998) Mersenne Twister used in EViews 4 (“mt4”). For parallel analysis only (“n=parallel”). | |
mfmethod=arg (default=“user”) | Maximum number of components used by selection methods: “schwert” (Schwert’s rule, default), “ah” (Ahn and Horenstein’s (2013) suggestion), “rootsize” ( 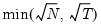 ), “size” ( ), “size” (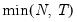 ), “user” (user specified value), where ), “user” (user specified value), where 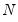 is the number of series and is the number of series and 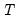 is the number of observations. is the number of observations.(1) For use with all components retention methods apart from user-specified (“fsmethod=user”). (2) If setting “mfmethod=user”, you may specify the maximum number of components using “rmax=”. (3) Schwert’s rule sets the maximum number of components using the rule: let 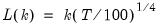 for 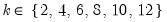 and let and let 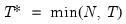 ; then the default maximum lag is given by ; then the default maximum lag is given by 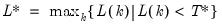 |
rmax=arg (default=all) | User-specified maximum number of factors to retain (for use when “mfmethod=user”). |
fsic=arg (default=avg) | Factor selection criterion (when “fsmethod=bn”): “icp1” (ICP1), “icp2” (ICP2), “icp3” (ICP3), “pcp1” (PCP1), “pcp2” (PCP1), “pcp3” (ICP3), “avg” (average of all criteria ICP1 through PCP3). Factor selection criterion (when “fsmethod=ah”): “er” (eigenvalue ratio), “gr” (growth ratio), “avg” (average of eigenvalue ratio and growth ratio). Factor selection criterion (when “fsmethod=simple”): “min” (minimum of: minimum eigenvalue, cumulative eigenvalue proportion, and maximum number of factors), “max” (maximum of: minimum eigenvalue, cumulative eigenvalue proportion, and maximum number of factors), “avg” (average the optimal number of factors as specified by the min and max rule, then round to the nearest integer). |
demeantime | Demeans observations across time prior to component selection procedures, when “n=bn” or “n=ah”. |
sdizetime | Standardizes observations across time prior to component selection procedures, when “n=bn” or “n=ah”. |
demeancross | Demeans observations across cross-sections prior to component selection procedures, when “n=bn” or “n=ah”. |
sdizecross | Standardizes observations across cross-sections prior to component selection procedures, when “n=bn” or “n=ah”. |